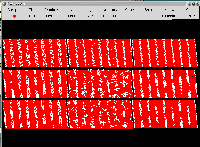
Nine MC simulations of atom deposition on a substrate, each one
with 44,000 atoms, to study the best conditions to achieve thin,
well defined tracks, corresponding to a total of 396,000 atoms,
all represented here as solid spheres. Each system is in a different,
transparent layer, with different observer coordinates, so the
atom coordinates are unchanged. To get the proper positioning,
each system was initially simulated by an orthorhombic cell
with the same size, replaced in the end by the system, editing
directly the file. Maura E. Monville, Oak Ridge National Laboratory.
Size: 80,265 bytes.
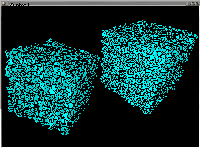
Voronoi tesselation, with periodic boundary conditions, of 10,000 atoms,
randomly positioned (on the left), and randomly positioned with a
minimum distance between them (4.05% of the cell width), corresponding
to a hard-sphere model (on the right). The Voronoi polyhedra obtained
are more smooth and regular for the restricted random (Laguerre)
than for the purely random (Poisson) distribution. A. Ferro,
N. Reis and C. Pereira, Technical University of Lisboa.
Size: 65,495 bytes.
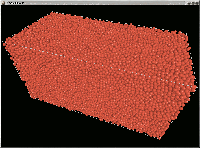
Orthorhombic cell containing 50,000 Cu atoms, forming a random
close packing (RCP) structure, with a density (occupied volume /
total volume) of 0.627. The structure was obtained using the
Jodrey algorithm with periodic boundary conditions. Increasing
the relaxation time, the density increases slightly (in the range
0.62-0.64, as found experimentaly in the random packing of large
numbers of equal spheres). Size: 171,181 bytes.
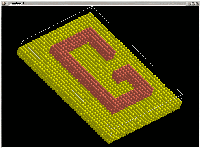
This multi-layer nanostructure was built linking 38x26x5 cP lattice
cells (a = 1.0) with substrate and adatoms (radius = 0.5). By
sucessively changing the cell origin, the GAMGI
occupancy rules can be used to create arbitrary, well defined,
blocks of atoms, forming structures such as the one seen in the picture
(see detailed procedure in the
tutorial).
Size: 78,354 bytes.
 Nine MC simulations of atom deposition on a substrate, each one
with 44,000 atoms, to study the best conditions to achieve thin,
well defined tracks, corresponding to a total of 396,000 atoms,
all represented here as solid spheres. Each system is in a different,
transparent layer, with different observer coordinates, so the
atom coordinates are unchanged. To get the proper positioning,
each system was initially simulated by an orthorhombic cell
with the same size, replaced in the end by the system, editing
directly the file. Maura E. Monville, Oak Ridge National Laboratory.
Size: 80,265 bytes.
Nine MC simulations of atom deposition on a substrate, each one
with 44,000 atoms, to study the best conditions to achieve thin,
well defined tracks, corresponding to a total of 396,000 atoms,
all represented here as solid spheres. Each system is in a different,
transparent layer, with different observer coordinates, so the
atom coordinates are unchanged. To get the proper positioning,
each system was initially simulated by an orthorhombic cell
with the same size, replaced in the end by the system, editing
directly the file. Maura E. Monville, Oak Ridge National Laboratory.
Size: 80,265 bytes.
 Voronoi tesselation, with periodic boundary conditions, of 10,000 atoms,
randomly positioned (on the left), and randomly positioned with a
minimum distance between them (4.05% of the cell width), corresponding
to a hard-sphere model (on the right). The Voronoi polyhedra obtained
are more smooth and regular for the restricted random (Laguerre)
than for the purely random (Poisson) distribution. A. Ferro,
N. Reis and C. Pereira, Technical University of Lisboa.
Size: 65,495 bytes.
Voronoi tesselation, with periodic boundary conditions, of 10,000 atoms,
randomly positioned (on the left), and randomly positioned with a
minimum distance between them (4.05% of the cell width), corresponding
to a hard-sphere model (on the right). The Voronoi polyhedra obtained
are more smooth and regular for the restricted random (Laguerre)
than for the purely random (Poisson) distribution. A. Ferro,
N. Reis and C. Pereira, Technical University of Lisboa.
Size: 65,495 bytes.
 Orthorhombic cell containing 50,000 Cu atoms, forming a random
close packing (RCP) structure, with a density (occupied volume /
total volume) of 0.627. The structure was obtained using the
Jodrey algorithm with periodic boundary conditions. Increasing
the relaxation time, the density increases slightly (in the range
0.62-0.64, as found experimentaly in the random packing of large
numbers of equal spheres). Size: 171,181 bytes.
Orthorhombic cell containing 50,000 Cu atoms, forming a random
close packing (RCP) structure, with a density (occupied volume /
total volume) of 0.627. The structure was obtained using the
Jodrey algorithm with periodic boundary conditions. Increasing
the relaxation time, the density increases slightly (in the range
0.62-0.64, as found experimentaly in the random packing of large
numbers of equal spheres). Size: 171,181 bytes.
 This multi-layer nanostructure was built linking 38x26x5 cP lattice
cells (a = 1.0) with substrate and adatoms (radius = 0.5). By
sucessively changing the cell origin, the GAMGI
occupancy rules can be used to create arbitrary, well defined,
blocks of atoms, forming structures such as the one seen in the picture
(see detailed procedure in the
tutorial).
Size: 78,354 bytes.
This multi-layer nanostructure was built linking 38x26x5 cP lattice
cells (a = 1.0) with substrate and adatoms (radius = 0.5). By
sucessively changing the cell origin, the GAMGI
occupancy rules can be used to create arbitrary, well defined,
blocks of atoms, forming structures such as the one seen in the picture
(see detailed procedure in the
tutorial).
Size: 78,354 bytes.